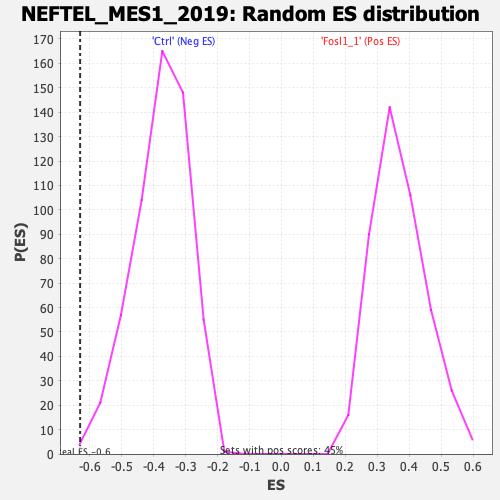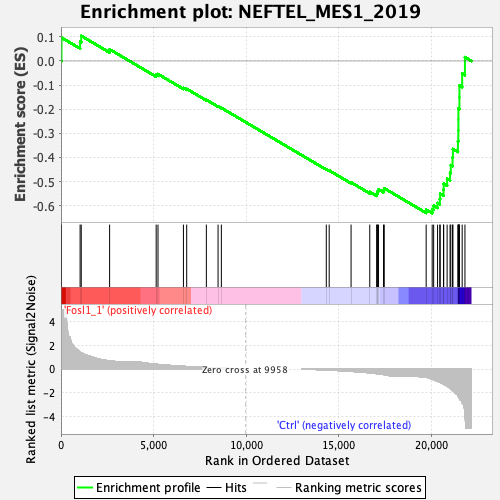
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | ranaseq_carol_ns_sel_exprs.ranaseq_carol_ns_sel_phenotype.cls #Fosl1_1_versus_Ctrl |
| Phenotype | ranaseq_carol_ns_sel_phenotype.cls#Fosl1_1_versus_Ctrl |
| Upregulated in class | Ctrl |
| GeneSet | NEFTEL_MES1_2019 |
| Enrichment Score (ES) | -0.6308056 |
| Normalized Enrichment Score (NES) | -1.6670926 |
| Nominal p-value | 0.0018018018 |
| FDR q-value | 0.001781849 |
| FWER p-Value | 0.024 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | SERPING1 | SERPING1 | 28 | 5.000 | 0.0969 | No |
| 2 | GFPT2 | GFPT2 | 1032 | 1.434 | 0.0798 | No |
| 3 | S100A16 | S100A16 | 1093 | 1.381 | 0.1042 | No |
| 4 | APOE | APOE | 2626 | 0.685 | 0.0484 | No |
| 5 | PRDX6 | PRDX6 | 5142 | 0.400 | -0.0573 | No |
| 6 | LGALS3 | LGALS3 | 5240 | 0.386 | -0.0542 | No |
| 7 | MGST1 | MGST1 | 6614 | 0.224 | -0.1118 | No |
| 8 | RCAN1 | RCAN1 | 6791 | 0.207 | -0.1157 | No |
| 9 | SOD2 | SOD2 | 7853 | 0.129 | -0.1611 | No |
| 10 | NPC2 | NPC2 | 8485 | 0.088 | -0.1878 | No |
| 11 | C3 | C3 | 8663 | 0.075 | -0.1944 | No |
| 12 | CTSB | CTSB | 14332 | -0.083 | -0.4488 | No |
| 13 | CLIC4 | CLIC4 | 14489 | -0.094 | -0.4540 | No |
| 14 | GJA1 | GJA1 | 15671 | -0.196 | -0.5035 | No |
| 15 | TUBA1C | TUBA1C | 16681 | -0.326 | -0.5427 | No |
| 16 | WWTR1 | WWTR1 | 17052 | -0.391 | -0.5517 | No |
| 17 | VIM | VIM | 17087 | -0.398 | -0.5454 | No |
| 18 | S100A10 | S100A10 | 17113 | -0.401 | -0.5387 | No |
| 19 | GSN | GSN | 17154 | -0.410 | -0.5324 | No |
| 20 | TNFRSF1A | TNFRSF1A | 17424 | -0.466 | -0.5354 | No |
| 21 | NAMPT | NAMPT | 17463 | -0.475 | -0.5278 | No |
| 22 | IGFBP7 | IGFBP7 | 19728 | -0.709 | -0.6162 | Yes |
| 23 | A2M | A2M | 20053 | -0.858 | -0.6140 | Yes |
| 24 | CLIC1 | CLIC1 | 20133 | -0.913 | -0.5996 | Yes |
| 25 | ACTN1 | ACTN1 | 20344 | -1.040 | -0.5887 | Yes |
| 26 | LGALS1 | LGALS1 | 20466 | -1.122 | -0.5721 | Yes |
| 27 | CD44 | CD44 | 20484 | -1.138 | -0.5505 | Yes |
| 28 | IFITM3 | IFITM3 | 20666 | -1.284 | -0.5335 | Yes |
| 29 | EMP1 | EMP1 | 20675 | -1.290 | -0.5085 | Yes |
| 30 | TAGLN2 | TAGLN2 | 20860 | -1.446 | -0.4884 | Yes |
| 31 | TIMP1 | TIMP1 | 21019 | -1.640 | -0.4633 | Yes |
| 32 | ANXA2 | ANXA2 | 21058 | -1.695 | -0.4318 | Yes |
| 33 | MGP | MGP | 21155 | -1.824 | -0.4003 | Yes |
| 34 | FN1 | FN1 | 21172 | -1.840 | -0.3649 | Yes |
| 35 | SERPINE1 | SERPINE1 | 21444 | -2.262 | -0.3327 | Yes |
| 36 | SPP1 | SPP1 | 21467 | -2.320 | -0.2882 | Yes |
| 37 | EMP3 | EMP3 | 21469 | -2.325 | -0.2426 | Yes |
| 38 | ANXA1 | ANXA1 | 21470 | -2.327 | -0.1969 | Yes |
| 39 | NNMT | NNMT | 21524 | -2.484 | -0.1505 | Yes |
| 40 | S100A11 | S100A11 | 21526 | -2.486 | -0.1017 | Yes |
| 41 | EFEMP1 | EFEMP1 | 21673 | -2.868 | -0.0520 | Yes |
| 42 | PLP2 | PLP2 | 21823 | -3.803 | 0.0159 | Yes |