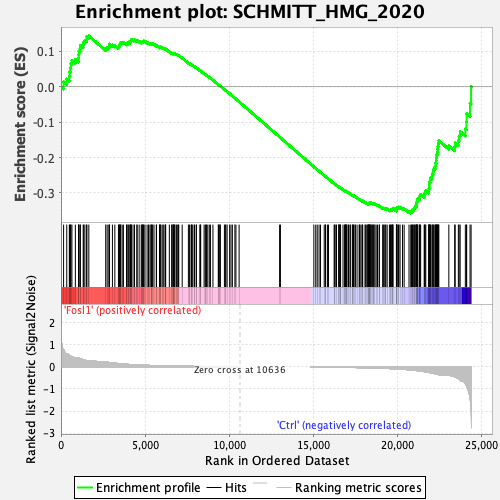
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NSCs_shNf1_sgFosl1.NSCs_shNf1_sgFosl1.cls #Fosl1_versus_Ctrl.NSCs_shNf1_sgFosl1.cls #Fosl1_versus_Ctrl_repos |
| Phenotype | NSCs_shNf1_sgFosl1.cls#Fosl1_versus_Ctrl_repos |
| Upregulated in class | Ctrl |
| GeneSet | SCHMITT_HMG_2020 |
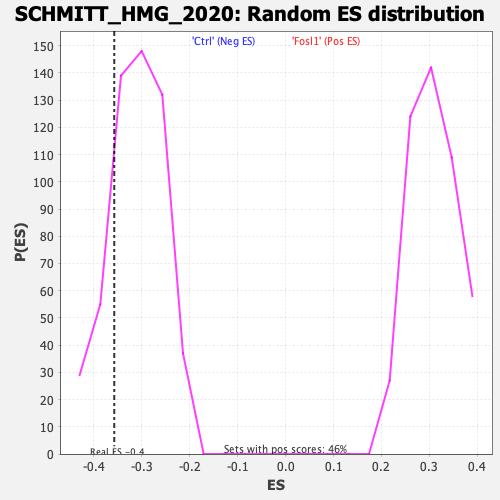
| Enrichment Score (ES) | -0.35737228 |
| Normalized Enrichment Score (NES) | -1.151571 |
| Nominal p-value | 0.18888889 |
| FDR q-value | 0.37685576 |
| FWER p-Value | 0.934 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | PMP2 | PMP2 | 154 | 0.771 | 0.0142 | No |
| 2 | PTGES | PTGES | 341 | 0.587 | 0.0221 | No |
| 3 | GRIK1 | GRIK1 | 494 | 0.526 | 0.0298 | No |
| 4 | ICAM1 | ICAM1 | 511 | 0.517 | 0.0429 | No |
| 5 | LBHD1 | LBHD1 | 563 | 0.487 | 0.0538 | No |
| 6 | JAK3 | JAK3 | 589 | 0.477 | 0.0655 | No |
| 7 | IL13RA1 | IL13RA1 | 649 | 0.452 | 0.0751 | No |
| 8 | GDA | GDA | 852 | 0.418 | 0.0778 | No |
| 9 | GPX3 | GPX3 | 1040 | 0.403 | 0.0808 | No |
| 10 | PDK4 | PDK4 | 1050 | 0.401 | 0.0912 | No |
| 11 | SLC16A4 | SLC16A4 | 1066 | 0.397 | 0.1011 | No |
| 12 | C1QL1 | C1QL1 | 1134 | 0.374 | 0.1083 | No |
| 13 | CCNB1IP1 | CCNB1IP1 | 1150 | 0.369 | 0.1175 | No |
| 14 | OSMR | OSMR | 1318 | 0.324 | 0.1192 | No |
| 15 | MYO5B | MYO5B | 1349 | 0.316 | 0.1264 | No |
| 16 | CCDC113 | CCDC113 | 1419 | 0.301 | 0.1315 | No |
| 17 | SOCS3 | SOCS3 | 1528 | 0.284 | 0.1346 | No |
| 18 | BCAM | BCAM | 1533 | 0.282 | 0.1420 | No |
| 19 | EFNA1 | EFNA1 | 1653 | 0.263 | 0.1441 | No |
| 20 | NDRG2 | NDRG2 | 2656 | 0.228 | 0.1086 | No |
| 21 | LAPTM5 | LAPTM5 | 2742 | 0.215 | 0.1108 | No |
| 22 | WIPI1 | WIPI1 | 2844 | 0.202 | 0.1120 | No |
| 23 | APOL6 | APOL6 | 2872 | 0.198 | 0.1162 | No |
| 24 | KCNK10 | KCNK10 | 2884 | 0.197 | 0.1210 | No |
| 25 | CLU | CLU | 3055 | 0.180 | 0.1187 | No |
| 26 | ALPK1 | ALPK1 | 3199 | 0.167 | 0.1172 | No |
| 27 | VASH2 | VASH2 | 3413 | 0.148 | 0.1123 | No |
| 28 | PTP4A3 | PTP4A3 | 3444 | 0.145 | 0.1150 | No |
| 29 | ZFP36 | ZFP36 | 3467 | 0.143 | 0.1179 | No |
| 30 | MEIS3 | MEIS3 | 3494 | 0.141 | 0.1205 | No |
| 31 | TIMP1 | TIMP1 | 3528 | 0.139 | 0.1229 | No |
| 32 | LIN7A | LIN7A | 3548 | 0.137 | 0.1257 | No |
| 33 | UBALD2 | UBALD2 | 3635 | 0.131 | 0.1257 | No |
| 34 | CBX7 | CBX7 | 3708 | 0.128 | 0.1261 | No |
| 35 | PROS1 | PROS1 | 3891 | 0.116 | 0.1216 | No |
| 36 | DUSP5 | DUSP5 | 3914 | 0.115 | 0.1238 | No |
| 37 | HOTAIRM1 | HOTAIRM1 | 3989 | 0.112 | 0.1237 | No |
| 38 | SPTB | SPTB | 3992 | 0.112 | 0.1266 | No |
| 39 | CISH | CISH | 4095 | 0.107 | 0.1252 | No |
| 40 | LIFR | LIFR | 4103 | 0.106 | 0.1277 | No |
| 41 | EEPD1 | EEPD1 | 4134 | 0.105 | 0.1293 | No |
| 42 | HSD3B7 | HSD3B7 | 4159 | 0.104 | 0.1311 | No |
| 43 | YPEL3 | YPEL3 | 4172 | 0.104 | 0.1334 | No |
| 44 | ZBTB46 | ZBTB46 | 4217 | 0.102 | 0.1342 | No |
| 45 | CIB1 | CIB1 | 4338 | 0.096 | 0.1318 | No |
| 46 | GLRX | GLRX | 4365 | 0.095 | 0.1333 | No |
| 47 | ZFP28 | ZFP28 | 4510 | 0.088 | 0.1297 | No |
| 48 | C4A | C4A | 4520 | 0.088 | 0.1316 | No |
| 49 | SLC2A4 | SLC2A4 | 4643 | 0.083 | 0.1288 | No |
| 50 | ENO3 | ENO3 | 4765 | 0.079 | 0.1259 | No |
| 51 | TCP11L2 | TCP11L2 | 4804 | 0.078 | 0.1264 | No |
| 52 | PORCN | PORCN | 4850 | 0.076 | 0.1265 | No |
| 53 | SRPX | SRPX | 4855 | 0.076 | 0.1284 | No |
| 54 | IFITM2 | IFITM2 | 4895 | 0.075 | 0.1288 | No |
| 55 | NRCAM | NRCAM | 4924 | 0.075 | 0.1296 | No |
| 56 | GADD45A | GADD45A | 4991 | 0.073 | 0.1288 | No |
| 57 | CITED1 | CITED1 | 5116 | 0.069 | 0.1255 | No |
| 58 | USP51 | USP51 | 5197 | 0.067 | 0.1240 | No |
| 59 | F3 | F3 | 5284 | 0.065 | 0.1222 | No |
| 60 | ARID5A | ARID5A | 5354 | 0.063 | 0.1210 | No |
| 61 | MCEE | MCEE | 5364 | 0.063 | 0.1223 | No |
| 62 | YPEL2 | YPEL2 | 5405 | 0.062 | 0.1223 | No |
| 63 | MVP | MVP | 5446 | 0.061 | 0.1223 | No |
| 64 | SH3BP5 | SH3BP5 | 5525 | 0.060 | 0.1206 | No |
| 65 | PPM1K | PPM1K | 5666 | 0.057 | 0.1163 | No |
| 66 | YIPF2 | YIPF2 | 5668 | 0.057 | 0.1178 | No |
| 67 | APOE | APOE | 5860 | 0.053 | 0.1113 | No |
| 68 | ARRDC3 | ARRDC3 | 5864 | 0.053 | 0.1126 | No |
| 69 | RPL37A | RPL37A | 5895 | 0.052 | 0.1127 | No |
| 70 | PIK3IP1 | PIK3IP1 | 5928 | 0.052 | 0.1128 | No |
| 71 | ABHD4 | ABHD4 | 6011 | 0.050 | 0.1107 | No |
| 72 | PLSCR4 | PLSCR4 | 6078 | 0.049 | 0.1093 | No |
| 73 | BCL6 | BCL6 | 6184 | 0.048 | 0.1062 | No |
| 74 | AGT | AGT | 6195 | 0.047 | 0.1070 | No |
| 75 | ADGRD1 | ADGRD1 | 6220 | 0.047 | 0.1073 | No |
| 76 | PITPNC1 | PITPNC1 | 6436 | 0.043 | 0.0995 | No |
| 77 | CIR1 | CIR1 | 6577 | 0.041 | 0.0948 | No |
| 78 | TAF1D | TAF1D | 6596 | 0.041 | 0.0952 | No |
| 79 | LGMN | LGMN | 6653 | 0.040 | 0.0939 | No |
| 80 | CRELD1 | CRELD1 | 6679 | 0.040 | 0.0940 | No |
| 81 | FGFBP3 | FGFBP3 | 6702 | 0.040 | 0.0941 | No |
| 82 | EIF2AK3 | EIF2AK3 | 6703 | 0.040 | 0.0952 | No |
| 83 | TNFRSF1A | TNFRSF1A | 6759 | 0.039 | 0.0939 | No |
| 84 | ARF4 | ARF4 | 6765 | 0.039 | 0.0947 | No |
| 85 | ACSL6 | ACSL6 | 6868 | 0.037 | 0.0915 | No |
| 86 | SNHG20 | SNHG20 | 6908 | 0.037 | 0.0909 | No |
| 87 | PIM1 | PIM1 | 6989 | 0.036 | 0.0885 | No |
| 88 | RAB4A | RAB4A | 6990 | 0.036 | 0.0894 | No |
| 89 | SC5D | SC5D | 7200 | 0.033 | 0.0817 | No |
| 90 | TMEM231 | TMEM231 | 7568 | 0.029 | 0.0672 | No |
| 91 | PLA2G4C | PLA2G4C | 7624 | 0.028 | 0.0657 | No |
| 92 | TPST1 | TPST1 | 7662 | 0.027 | 0.0649 | No |
| 93 | CYB561D2 | CYB561D2 | 7756 | 0.027 | 0.0617 | No |
| 94 | YOD1 | YOD1 | 7758 | 0.027 | 0.0624 | No |
| 95 | ATP1B2 | ATP1B2 | 7802 | 0.026 | 0.0613 | No |
| 96 | POLR3D | POLR3D | 7906 | 0.025 | 0.0577 | No |
| 97 | CSF1 | CSF1 | 7934 | 0.025 | 0.0572 | No |
| 98 | TMEM97 | TMEM97 | 8038 | 0.023 | 0.0536 | No |
| 99 | MOSPD1 | MOSPD1 | 8040 | 0.023 | 0.0542 | No |
| 100 | DRAM2 | DRAM2 | 8249 | 0.021 | 0.0461 | No |
| 101 | SLC50A1 | SLC50A1 | 8282 | 0.021 | 0.0453 | No |
| 102 | CBS | CBS | 8300 | 0.021 | 0.0452 | No |
| 103 | PFKFB2 | PFKFB2 | 8502 | 0.019 | 0.0373 | No |
| 104 | SYN3 | SYN3 | 8573 | 0.018 | 0.0349 | No |
| 105 | THBS2 | THBS2 | 8609 | 0.018 | 0.0339 | No |
| 106 | PTCHD1 | PTCHD1 | 8641 | 0.017 | 0.0331 | No |
| 107 | RGS3 | RGS3 | 8720 | 0.017 | 0.0303 | No |
| 108 | TMEM50B | TMEM50B | 8811 | 0.016 | 0.0270 | No |
| 109 | S100A10 | S100A10 | 8879 | 0.015 | 0.0247 | No |
| 110 | CLIP2 | CLIP2 | 9027 | 0.014 | 0.0189 | No |
| 111 | VMAC | VMAC | 9344 | 0.011 | 0.0061 | No |
| 112 | TAGLN2 | TAGLN2 | 9367 | 0.011 | 0.0055 | No |
| 113 | AKNA | AKNA | 9394 | 0.011 | 0.0047 | No |
| 114 | STK19 | STK19 | 9439 | 0.010 | 0.0032 | No |
| 115 | MAP3K6 | MAP3K6 | 9469 | 0.010 | 0.0022 | No |
| 116 | CRTAP | CRTAP | 9709 | 0.008 | -0.0075 | No |
| 117 | ATCAY | ATCAY | 9761 | 0.007 | -0.0094 | No |
| 118 | DDR1 | DDR1 | 9793 | 0.007 | -0.0105 | No |
| 119 | PIKFYVE | PIKFYVE | 9869 | 0.007 | -0.0134 | No |
| 120 | TBC1D23 | TBC1D23 | 10019 | 0.005 | -0.0195 | No |
| 121 | CDHR1 | CDHR1 | 10099 | 0.005 | -0.0226 | No |
| 122 | GOLT1B | GOLT1B | 10182 | 0.004 | -0.0259 | No |
| 123 | PMP22 | PMP22 | 10332 | 0.003 | -0.0320 | No |
| 124 | MYCN | MYCN | 10393 | 0.002 | -0.0344 | No |
| 125 | N4BP2L1 | N4BP2L1 | 10582 | 0.000 | -0.0422 | No |
| 126 | HMGCS1 | HMGCS1 | 12988 | 0.000 | -0.1420 | No |
| 127 | IL21R | IL21R | 13037 | 0.000 | -0.1440 | No |
| 128 | BLOC1S3 | BLOC1S3 | 15023 | -0.002 | -0.2262 | No |
| 129 | UFM1 | UFM1 | 15136 | -0.003 | -0.2308 | No |
| 130 | COG6 | COG6 | 15223 | -0.004 | -0.2343 | No |
| 131 | IDI1 | IDI1 | 15332 | -0.004 | -0.2386 | No |
| 132 | ACBD3 | ACBD3 | 15419 | -0.005 | -0.2421 | No |
| 133 | TSC22D3 | TSC22D3 | 15639 | -0.007 | -0.2510 | No |
| 134 | ZFPL1 | ZFPL1 | 15712 | -0.008 | -0.2537 | No |
| 135 | ACSS2 | ACSS2 | 15721 | -0.008 | -0.2538 | No |
| 136 | SERPINE1 | SERPINE1 | 15830 | -0.009 | -0.2581 | No |
| 137 | RAB3B | RAB3B | 15894 | -0.009 | -0.2605 | No |
| 138 | INSIG1 | INSIG1 | 16227 | -0.013 | -0.2739 | No |
| 139 | GOLGA2 | GOLGA2 | 16275 | -0.013 | -0.2755 | No |
| 140 | ERLEC1 | ERLEC1 | 16278 | -0.013 | -0.2752 | No |
| 141 | GPR107 | GPR107 | 16368 | -0.014 | -0.2785 | No |
| 142 | KDELR3 | KDELR3 | 16508 | -0.016 | -0.2839 | No |
| 143 | SLC30A5 | SLC30A5 | 16516 | -0.016 | -0.2837 | No |
| 144 | TMCC3 | TMCC3 | 16520 | -0.016 | -0.2834 | No |
| 145 | TMEM87B | TMEM87B | 16580 | -0.016 | -0.2854 | No |
| 146 | SEMA4B | SEMA4B | 16590 | -0.016 | -0.2854 | No |
| 147 | GOSR2 | GOSR2 | 16610 | -0.017 | -0.2857 | No |
| 148 | GOLGA4 | GOLGA4 | 16756 | -0.018 | -0.2913 | No |
| 149 | EPHA5 | EPHA5 | 16857 | -0.019 | -0.2949 | No |
| 150 | GOLGB1 | GOLGB1 | 16858 | -0.019 | -0.2944 | No |
| 151 | LMAN1 | LMAN1 | 16911 | -0.020 | -0.2960 | No |
| 152 | RAB5A | RAB5A | 16927 | -0.020 | -0.2961 | No |
| 153 | RDH10 | RDH10 | 16938 | -0.020 | -0.2959 | No |
| 154 | ZBTB41 | ZBTB41 | 16939 | -0.020 | -0.2954 | No |
| 155 | TRIM7 | TRIM7 | 16941 | -0.020 | -0.2949 | No |
| 156 | PDP2 | PDP2 | 17049 | -0.022 | -0.2988 | No |
| 157 | MSMO1 | MSMO1 | 17081 | -0.022 | -0.2995 | No |
| 158 | LDLR | LDLR | 17170 | -0.023 | -0.3025 | No |
| 159 | FBXO32 | FBXO32 | 17191 | -0.023 | -0.3027 | No |
| 160 | MATN2 | MATN2 | 17308 | -0.025 | -0.3068 | No |
| 161 | TBC1D15 | TBC1D15 | 17342 | -0.025 | -0.3075 | No |
| 162 | PDXDC1 | PDXDC1 | 17348 | -0.025 | -0.3071 | No |
| 163 | AOC2 | AOC2 | 17383 | -0.026 | -0.3078 | No |
| 164 | VMP1 | VMP1 | 17464 | -0.027 | -0.3104 | No |
| 165 | ITGA10 | ITGA10 | 17488 | -0.027 | -0.3106 | No |
| 166 | MTUS1 | MTUS1 | 17590 | -0.028 | -0.3141 | No |
| 167 | SGK1 | SGK1 | 17712 | -0.030 | -0.3183 | No |
| 168 | COL11A2 | COL11A2 | 17740 | -0.031 | -0.3186 | No |
| 169 | NPC2 | NPC2 | 17773 | -0.031 | -0.3191 | No |
| 170 | HABP4 | HABP4 | 17879 | -0.033 | -0.3226 | No |
| 171 | HMGXB3 | HMGXB3 | 17902 | -0.033 | -0.3226 | No |
| 172 | TMEM173 | TMEM173 | 17927 | -0.033 | -0.3227 | No |
| 173 | SYDE2 | SYDE2 | 18061 | -0.036 | -0.3273 | No |
| 174 | GOLGA5 | GOLGA5 | 18102 | -0.036 | -0.3280 | No |
| 175 | MALT1 | MALT1 | 18169 | -0.037 | -0.3297 | No |
| 176 | PPP2R5A | PPP2R5A | 18222 | -0.038 | -0.3309 | No |
| 177 | GCNT4 | GCNT4 | 18243 | -0.039 | -0.3307 | No |
| 178 | PMM2 | PMM2 | 18276 | -0.039 | -0.3309 | No |
| 179 | PJA2 | PJA2 | 18298 | -0.040 | -0.3307 | No |
| 180 | MAFF | MAFF | 18299 | -0.040 | -0.3297 | No |
| 181 | EDEM3 | EDEM3 | 18308 | -0.040 | -0.3290 | No |
| 182 | LONRF1 | LONRF1 | 18320 | -0.040 | -0.3283 | No |
| 183 | SNX10 | SNX10 | 18335 | -0.040 | -0.3278 | No |
| 184 | FAM114A1 | FAM114A1 | 18380 | -0.041 | -0.3286 | No |
| 185 | TRAF1 | TRAF1 | 18411 | -0.042 | -0.3287 | No |
| 186 | IFITM1 | IFITM1 | 18435 | -0.042 | -0.3285 | No |
| 187 | PDE4DIP | PDE4DIP | 18497 | -0.043 | -0.3299 | No |
| 188 | SEC14L1 | SEC14L1 | 18540 | -0.044 | -0.3305 | No |
| 189 | ACAN | ACAN | 18550 | -0.045 | -0.3296 | No |
| 190 | GOLGA3 | GOLGA3 | 18596 | -0.046 | -0.3303 | No |
| 191 | TMEM120A | TMEM120A | 18652 | -0.047 | -0.3313 | No |
| 192 | FMN2 | FMN2 | 18725 | -0.048 | -0.3330 | No |
| 193 | COL6A1 | COL6A1 | 18805 | -0.050 | -0.3350 | No |
| 194 | MAN2A1 | MAN2A1 | 18907 | -0.053 | -0.3377 | No |
| 195 | CHN2 | CHN2 | 18912 | -0.053 | -0.3365 | No |
| 196 | JAG2 | JAG2 | 19112 | -0.059 | -0.3432 | No |
| 197 | NUAK2 | NUAK2 | 19150 | -0.060 | -0.3431 | No |
| 198 | RUNX1T1 | RUNX1T1 | 19202 | -0.062 | -0.3436 | No |
| 199 | CCNG1 | CCNG1 | 19260 | -0.063 | -0.3443 | No |
| 200 | DNAJB5 | DNAJB5 | 19300 | -0.065 | -0.3442 | No |
| 201 | CDKL3 | CDKL3 | 19378 | -0.067 | -0.3456 | No |
| 202 | KCNA4 | KCNA4 | 19512 | -0.072 | -0.3492 | No |
| 203 | TXNIP | TXNIP | 19557 | -0.074 | -0.3490 | No |
| 204 | GJA3 | GJA3 | 19579 | -0.074 | -0.3479 | No |
| 205 | NAMPT | NAMPT | 19644 | -0.077 | -0.3485 | No |
| 206 | DBP | DBP | 19676 | -0.079 | -0.3477 | No |
| 207 | MKNK1 | MKNK1 | 19689 | -0.079 | -0.3461 | No |
| 208 | CDKN1A | CDKN1A | 19702 | -0.080 | -0.3445 | No |
| 209 | KLF9 | KLF9 | 19731 | -0.081 | -0.3435 | No |
| 210 | KLF6 | KLF6 | 19927 | -0.090 | -0.3492 | No |
| 211 | ARMCX1 | ARMCX1 | 19940 | -0.091 | -0.3472 | No |
| 212 | SPTY2D1 | SPTY2D1 | 19946 | -0.091 | -0.3450 | No |
| 213 | CD44 | CD44 | 19954 | -0.091 | -0.3429 | No |
| 214 | TNC | TNC | 19967 | -0.092 | -0.3409 | No |
| 215 | SLC26A2 | SLC26A2 | 20050 | -0.096 | -0.3418 | No |
| 216 | PODXL | PODXL | 20071 | -0.097 | -0.3400 | No |
| 217 | PDP1 | PDP1 | 20145 | -0.101 | -0.3404 | No |
| 218 | CLDN4 | CLDN4 | 20291 | -0.107 | -0.3435 | No |
| 219 | HDAC9 | HDAC9 | 20388 | -0.111 | -0.3445 | No |
| 220 | DGKI | DGKI | 20677 | -0.127 | -0.3531 | No |
| 221 | EPHX1 | EPHX1 | 20781 | -0.132 | -0.3538 | Yes |
| 222 | FZD5 | FZD5 | 20828 | -0.135 | -0.3521 | Yes |
| 223 | RPE65 | RPE65 | 20877 | -0.138 | -0.3504 | Yes |
| 224 | JADE2 | JADE2 | 20889 | -0.140 | -0.3472 | Yes |
| 225 | GLIS3 | GLIS3 | 20957 | -0.147 | -0.3460 | Yes |
| 226 | IRF9 | IRF9 | 20986 | -0.150 | -0.3432 | Yes |
| 227 | CAMK2D | CAMK2D | 21027 | -0.154 | -0.3407 | Yes |
| 228 | DTX3L | DTX3L | 21046 | -0.156 | -0.3373 | Yes |
| 229 | SPARC | SPARC | 21114 | -0.162 | -0.3358 | Yes |
| 230 | ABCD2 | ABCD2 | 21118 | -0.163 | -0.3316 | Yes |
| 231 | GPR20 | GPR20 | 21135 | -0.164 | -0.3278 | Yes |
| 232 | PKIG | PKIG | 21151 | -0.165 | -0.3241 | Yes |
| 233 | NEDD9 | NEDD9 | 21158 | -0.166 | -0.3199 | Yes |
| 234 | EMP1 | EMP1 | 21196 | -0.170 | -0.3169 | Yes |
| 235 | FUT1 | FUT1 | 21286 | -0.180 | -0.3158 | Yes |
| 236 | NUPR1 | NUPR1 | 21314 | -0.184 | -0.3120 | Yes |
| 237 | GBP2 | GBP2 | 21326 | -0.185 | -0.3075 | Yes |
| 238 | ATF3 | ATF3 | 21368 | -0.190 | -0.3041 | Yes |
| 239 | ANXA11 | ANXA11 | 21579 | -0.211 | -0.3072 | Yes |
| 240 | SLC22A18 | SLC22A18 | 21596 | -0.212 | -0.3022 | Yes |
| 241 | BST2 | BST2 | 21635 | -0.216 | -0.2981 | Yes |
| 242 | RCN3 | RCN3 | 21673 | -0.220 | -0.2937 | Yes |
| 243 | BMP5 | BMP5 | 21844 | -0.245 | -0.2942 | Yes |
| 244 | PRICKLE2 | PRICKLE2 | 21857 | -0.248 | -0.2881 | Yes |
| 245 | A2M | A2M | 21880 | -0.251 | -0.2823 | Yes |
| 246 | EML2 | EML2 | 21882 | -0.251 | -0.2757 | Yes |
| 247 | CRYAB | CRYAB | 21884 | -0.252 | -0.2690 | Yes |
| 248 | TRIQK | TRIQK | 21944 | -0.260 | -0.2645 | Yes |
| 249 | SYNM | SYNM | 21947 | -0.261 | -0.2577 | Yes |
| 250 | LYST | LYST | 22043 | -0.276 | -0.2542 | Yes |
| 251 | SLC2A3 | SLC2A3 | 22057 | -0.278 | -0.2474 | Yes |
| 252 | LIPG | LIPG | 22107 | -0.286 | -0.2418 | Yes |
| 253 | COL6A2 | COL6A2 | 22117 | -0.287 | -0.2345 | Yes |
| 254 | VGF | VGF | 22175 | -0.298 | -0.2289 | Yes |
| 255 | TLCD2 | TLCD2 | 22251 | -0.311 | -0.2237 | Yes |
| 256 | TTC39B | TTC39B | 22254 | -0.312 | -0.2155 | Yes |
| 257 | FGD4 | FGD4 | 22299 | -0.321 | -0.2088 | Yes |
| 258 | NR4A3 | NR4A3 | 22302 | -0.321 | -0.2003 | Yes |
| 259 | SYCP2 | SYCP2 | 22322 | -0.326 | -0.1924 | Yes |
| 260 | ATG9B | ATG9B | 22362 | -0.333 | -0.1851 | Yes |
| 261 | MERTK | MERTK | 22373 | -0.334 | -0.1766 | Yes |
| 262 | FABP3 | FABP3 | 22389 | -0.337 | -0.1682 | Yes |
| 263 | COL14A1 | COL14A1 | 22421 | -0.344 | -0.1603 | Yes |
| 264 | TRPM8 | TRPM8 | 22449 | -0.349 | -0.1522 | Yes |
| 265 | FSTL5 | FSTL5 | 23043 | -0.372 | -0.1668 | Yes |
| 266 | GREM2 | GREM2 | 23392 | -0.443 | -0.1695 | Yes |
| 267 | RGS16 | RGS16 | 23418 | -0.454 | -0.1584 | Yes |
| 268 | AHNAK | AHNAK | 23609 | -0.531 | -0.1521 | Yes |
| 269 | PGM2L1 | PGM2L1 | 23652 | -0.545 | -0.1393 | Yes |
| 270 | PHF24 | PHF24 | 23716 | -0.585 | -0.1264 | Yes |
| 271 | ISG20 | ISG20 | 24023 | -0.758 | -0.1188 | Yes |
| 272 | PDGFRB | PDGFRB | 24091 | -0.852 | -0.0989 | Yes |
| 273 | THEMIS2 | THEMIS2 | 24103 | -0.877 | -0.0760 | Yes |
| 274 | ITGA11 | ITGA11 | 24303 | -1.408 | -0.0467 | Yes |
| 275 | LHX9 | LHX9 | 24365 | -1.883 | 0.0010 | Yes |