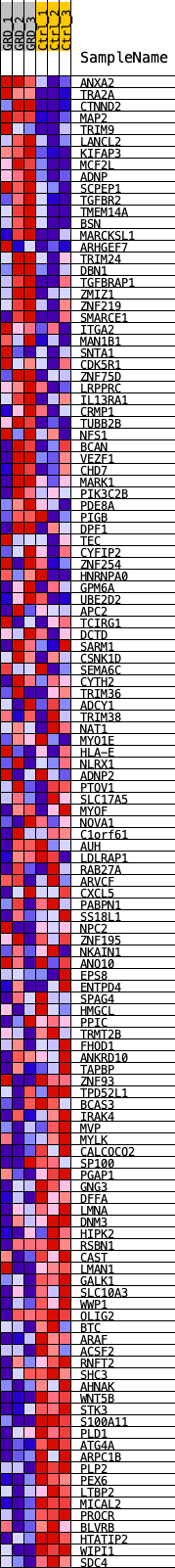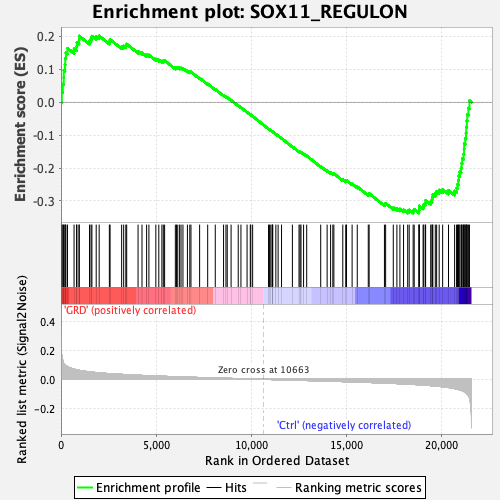
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NF1_GRD_exprs.NF1_GRD_phenotype.cls #GRD_versus_Ctrl.NF1_GRD_phenotype.cls #GRD_versus_Ctrl_repos |
| Phenotype | NF1_GRD_phenotype.cls#GRD_versus_Ctrl_repos |
| Upregulated in class | Ctrl |
| GeneSet | SOX11_REGULON |
| Enrichment Score (ES) | -0.33689862 |
| Normalized Enrichment Score (NES) | -1.4327704 |
| Nominal p-value | 0.015779093 |
| FDR q-value | 0.031647433 |
| FWER p-Value | 0.305 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | ANXA2 | ANXA2 | 41 | 0.164 | 0.0313 | No |
| 2 | TRA2A | TRA2A | 95 | 0.129 | 0.0549 | No |
| 3 | CTNND2 | CTNND2 | 142 | 0.109 | 0.0750 | No |
| 4 | MAP2 | MAP2 | 145 | 0.108 | 0.0968 | No |
| 5 | TRIM9 | TRIM9 | 210 | 0.099 | 0.1140 | No |
| 6 | LANCL2 | LANCL2 | 224 | 0.096 | 0.1330 | No |
| 7 | KIFAP3 | KIFAP3 | 259 | 0.093 | 0.1502 | No |
| 8 | MCF2L | MCF2L | 343 | 0.087 | 0.1639 | No |
| 9 | ADNP | ADNP | 687 | 0.070 | 0.1622 | No |
| 10 | SCPEP1 | SCPEP1 | 820 | 0.065 | 0.1692 | No |
| 11 | TGFBR2 | TGFBR2 | 835 | 0.065 | 0.1817 | No |
| 12 | TMEM14A | TMEM14A | 950 | 0.062 | 0.1889 | No |
| 13 | BSN | BSN | 956 | 0.062 | 0.2012 | No |
| 14 | MARCKSL1 | MARCKSL1 | 1500 | 0.051 | 0.1863 | No |
| 15 | ARHGEF7 | ARHGEF7 | 1579 | 0.050 | 0.1928 | No |
| 16 | TRIM24 | TRIM24 | 1626 | 0.049 | 0.2007 | No |
| 17 | DBN1 | DBN1 | 1852 | 0.046 | 0.1997 | No |
| 18 | TGFBRAP1 | TGFBRAP1 | 2008 | 0.045 | 0.2015 | No |
| 19 | ZMIZ1 | ZMIZ1 | 2544 | 0.039 | 0.1845 | No |
| 20 | ZNF219 | ZNF219 | 2582 | 0.039 | 0.1906 | No |
| 21 | SMARCE1 | SMARCE1 | 3190 | 0.034 | 0.1692 | No |
| 22 | ITGA2 | ITGA2 | 3297 | 0.033 | 0.1710 | No |
| 23 | MAN1B1 | MAN1B1 | 3411 | 0.032 | 0.1722 | No |
| 24 | SNTA1 | SNTA1 | 3456 | 0.032 | 0.1766 | No |
| 25 | CDK5R1 | CDK5R1 | 4054 | 0.028 | 0.1545 | No |
| 26 | ZNF75D | ZNF75D | 4261 | 0.027 | 0.1503 | No |
| 27 | LRPPRC | LRPPRC | 4504 | 0.025 | 0.1442 | No |
| 28 | IL13RA1 | IL13RA1 | 4625 | 0.025 | 0.1436 | No |
| 29 | CRMP1 | CRMP1 | 4989 | 0.023 | 0.1312 | No |
| 30 | TUBB2B | TUBB2B | 5145 | 0.022 | 0.1284 | No |
| 31 | NFS1 | NFS1 | 5308 | 0.021 | 0.1251 | No |
| 32 | BCAN | BCAN | 5383 | 0.021 | 0.1259 | No |
| 33 | VEZF1 | VEZF1 | 5451 | 0.020 | 0.1269 | No |
| 34 | CHD7 | CHD7 | 6015 | 0.018 | 0.1042 | No |
| 35 | MARK1 | MARK1 | 6078 | 0.017 | 0.1049 | No |
| 36 | PIK3C2B | PIK3C2B | 6142 | 0.017 | 0.1054 | No |
| 37 | PDE8A | PDE8A | 6241 | 0.017 | 0.1042 | No |
| 38 | PIGB | PIGB | 6315 | 0.016 | 0.1042 | No |
| 39 | DPF1 | DPF1 | 6416 | 0.016 | 0.1027 | No |
| 40 | TEC | TEC | 6651 | 0.015 | 0.0949 | No |
| 41 | CYFIP2 | CYFIP2 | 6776 | 0.014 | 0.0920 | No |
| 42 | ZNF254 | ZNF254 | 6837 | 0.014 | 0.0921 | No |
| 43 | HNRNPA0 | HNRNPA0 | 7293 | 0.012 | 0.0734 | No |
| 44 | GPM6A | GPM6A | 7717 | 0.011 | 0.0558 | No |
| 45 | UBE2D2 | UBE2D2 | 8116 | 0.009 | 0.0392 | No |
| 46 | APC2 | APC2 | 8552 | 0.008 | 0.0205 | No |
| 47 | TCIRG1 | TCIRG1 | 8678 | 0.007 | 0.0161 | No |
| 48 | DCTD | DCTD | 8745 | 0.007 | 0.0144 | No |
| 49 | SARM1 | SARM1 | 8948 | 0.006 | 0.0062 | No |
| 50 | CSNK1D | CSNK1D | 9324 | 0.005 | -0.0102 | No |
| 51 | SEMA6C | SEMA6C | 9467 | 0.004 | -0.0160 | No |
| 52 | CYTH2 | CYTH2 | 9789 | 0.003 | -0.0303 | No |
| 53 | TRIM36 | TRIM36 | 9958 | 0.002 | -0.0377 | No |
| 54 | ADCY1 | ADCY1 | 9986 | 0.002 | -0.0384 | No |
| 55 | TRIM38 | TRIM38 | 10082 | 0.002 | -0.0424 | No |
| 56 | NAT1 | NAT1 | 10918 | -0.001 | -0.0812 | No |
| 57 | MYO1E | MYO1E | 10966 | -0.001 | -0.0831 | No |
| 58 | HLA-E | HLA-E | 11037 | -0.001 | -0.0861 | No |
| 59 | NLRX1 | NLRX1 | 11106 | -0.002 | -0.0890 | No |
| 60 | ADNP2 | ADNP2 | 11127 | -0.002 | -0.0896 | No |
| 61 | PTOV1 | PTOV1 | 11141 | -0.002 | -0.0899 | No |
| 62 | SLC17A5 | SLC17A5 | 11303 | -0.002 | -0.0969 | No |
| 63 | MYOF | MYOF | 11422 | -0.003 | -0.1019 | No |
| 64 | NOVA1 | NOVA1 | 11602 | -0.003 | -0.1095 | No |
| 65 | C1orf61 | C1orf61 | 12173 | -0.005 | -0.1350 | No |
| 66 | AUH | AUH | 12522 | -0.007 | -0.1499 | No |
| 67 | LDLRAP1 | LDLRAP1 | 12576 | -0.007 | -0.1510 | No |
| 68 | RAB27A | RAB27A | 12626 | -0.007 | -0.1519 | No |
| 69 | ARVCF | ARVCF | 12762 | -0.007 | -0.1567 | No |
| 70 | CXCL5 | CXCL5 | 12926 | -0.008 | -0.1627 | No |
| 71 | PABPN1 | PABPN1 | 13663 | -0.011 | -0.1948 | No |
| 72 | SS18L1 | SS18L1 | 14001 | -0.012 | -0.2081 | No |
| 73 | NPC2 | NPC2 | 14183 | -0.013 | -0.2139 | No |
| 74 | ZNF195 | ZNF195 | 14301 | -0.013 | -0.2167 | No |
| 75 | NKAIN1 | NKAIN1 | 14356 | -0.013 | -0.2165 | No |
| 76 | ANO10 | ANO10 | 14825 | -0.015 | -0.2351 | No |
| 77 | EPS8 | EPS8 | 14980 | -0.016 | -0.2391 | No |
| 78 | ENTPD4 | ENTPD4 | 15021 | -0.016 | -0.2377 | No |
| 79 | SPAG4 | SPAG4 | 15315 | -0.017 | -0.2478 | No |
| 80 | HMGCL | HMGCL | 15586 | -0.019 | -0.2566 | No |
| 81 | PPIC | PPIC | 16165 | -0.021 | -0.2792 | No |
| 82 | TRMT2B | TRMT2B | 16211 | -0.021 | -0.2770 | No |
| 83 | FHOD1 | FHOD1 | 17016 | -0.025 | -0.3093 | No |
| 84 | ANKRD10 | ANKRD10 | 17079 | -0.026 | -0.3069 | No |
| 85 | TAPBP | TAPBP | 17481 | -0.028 | -0.3200 | No |
| 86 | ZNF93 | ZNF93 | 17671 | -0.029 | -0.3229 | No |
| 87 | TPD52L1 | TPD52L1 | 17830 | -0.030 | -0.3242 | No |
| 88 | BCAS3 | BCAS3 | 18021 | -0.031 | -0.3267 | No |
| 89 | IRAK4 | IRAK4 | 18240 | -0.033 | -0.3303 | Yes |
| 90 | MVP | MVP | 18318 | -0.033 | -0.3272 | Yes |
| 91 | MYLK | MYLK | 18526 | -0.035 | -0.3298 | Yes |
| 92 | CALCOCO2 | CALCOCO2 | 18581 | -0.035 | -0.3252 | Yes |
| 93 | SP100 | SP100 | 18830 | -0.037 | -0.3292 | Yes |
| 94 | PGAP1 | PGAP1 | 18844 | -0.037 | -0.3222 | Yes |
| 95 | GNG3 | GNG3 | 18848 | -0.037 | -0.3148 | Yes |
| 96 | DFFA | DFFA | 19050 | -0.039 | -0.3162 | Yes |
| 97 | LMNA | LMNA | 19085 | -0.039 | -0.3098 | Yes |
| 98 | DNM3 | DNM3 | 19167 | -0.040 | -0.3054 | Yes |
| 99 | HIPK2 | HIPK2 | 19182 | -0.040 | -0.2978 | Yes |
| 100 | RSBN1 | RSBN1 | 19450 | -0.043 | -0.3015 | Yes |
| 101 | CAST | CAST | 19519 | -0.044 | -0.2958 | Yes |
| 102 | LMAN1 | LMAN1 | 19532 | -0.044 | -0.2875 | Yes |
| 103 | GALK1 | GALK1 | 19562 | -0.044 | -0.2798 | Yes |
| 104 | SLC10A3 | SLC10A3 | 19691 | -0.046 | -0.2765 | Yes |
| 105 | WWP1 | WWP1 | 19762 | -0.047 | -0.2703 | Yes |
| 106 | OLIG2 | OLIG2 | 19896 | -0.049 | -0.2667 | Yes |
| 107 | BTC | BTC | 20078 | -0.051 | -0.2647 | Yes |
| 108 | ARAF | ARAF | 20386 | -0.056 | -0.2677 | Yes |
| 109 | ACSF2 | ACSF2 | 20708 | -0.064 | -0.2697 | Yes |
| 110 | RNFT2 | RNFT2 | 20811 | -0.067 | -0.2609 | Yes |
| 111 | SHC3 | SHC3 | 20860 | -0.068 | -0.2494 | Yes |
| 112 | AHNAK | AHNAK | 20906 | -0.070 | -0.2373 | Yes |
| 113 | WNT5B | WNT5B | 20925 | -0.070 | -0.2239 | Yes |
| 114 | STK3 | STK3 | 20977 | -0.072 | -0.2117 | Yes |
| 115 | S100A11 | S100A11 | 21047 | -0.075 | -0.1998 | Yes |
| 116 | PLD1 | PLD1 | 21074 | -0.076 | -0.1856 | Yes |
| 117 | ATG4A | ATG4A | 21108 | -0.078 | -0.1714 | Yes |
| 118 | ARPC1B | ARPC1B | 21177 | -0.082 | -0.1580 | Yes |
| 119 | PLP2 | PLP2 | 21203 | -0.083 | -0.1422 | Yes |
| 120 | PEX6 | PEX6 | 21211 | -0.084 | -0.1254 | Yes |
| 121 | LTBP2 | LTBP2 | 21266 | -0.088 | -0.1101 | Yes |
| 122 | MICAL2 | MICAL2 | 21308 | -0.091 | -0.0934 | Yes |
| 123 | PROCR | PROCR | 21321 | -0.093 | -0.0750 | Yes |
| 124 | BLVRB | BLVRB | 21350 | -0.098 | -0.0565 | Yes |
| 125 | HTATIP2 | HTATIP2 | 21373 | -0.101 | -0.0371 | Yes |
| 126 | WIPI1 | WIPI1 | 21433 | -0.109 | -0.0178 | Yes |
| 127 | SDC4 | SDC4 | 21489 | -0.125 | 0.0049 | Yes |